Haupt: Paper3
{{template:paper_2figures |abstract=
The objective of this research is to generate grain boundary structures over a wide range of degrees of freedom for future use in assessing how grain boundary degrees of freedom impact the properties of polycrystalline materials. For each grain boundary, this work uses a parallel molecular dynamics code, LAMMPS, with in-plane translations and atom deletion criteria to sample a large number of potential structures to find the global minimum energy grain boundary structure[1][2]. The results of this work are grain boundary structures that agree with previously calculated structures as well as experimentally-imaged HRTEM structures. Additionally, the grain boundary energies agree with previous calculated and measured energies. The significance of this research is that grain boundary properties play an important role in the properties of polycrystalline materials and this research enables future atomistic research investigating these properties.
|authors= Mark A. Tschopp, Mark F. Horstemeyer
Corresponding Author: Mark Tschopp
|image1= bicrystal_sim_cell.jpg
|image1 caption= Figure 1. An example of the bicrystal simulation cell used for the grain boundary structure calculations. In this particular example, the grain boundary is a <110> tilt grain boundary.
|image2= plot_accessibility.jpg
|image2 caption= Figure 2. This is a plot of the accessibility of each grain boundary structure for an asymmetric tilt grain boundary. For complex boundaries, a large number of boundaries may need to be sampled to find the "global" minimum energy boundary.
|methodology= For each grain boundary structure, the crystal orientations and their relationship to the grain boundary plane -- five degrees of freedom -- are often used to describe the grain boundary character. A three-dimensional periodic simulation cell is used for these calculations (see Fig. 1), whereby there are two crystal orientations and two grain boundaries (one in the middle and one at the top/bottom). This simulation cell allows the user to manipulate the five grain boundary degrees of freedom to investigate their relationship on properties. The distance between the two boundaries must be large enough for the grain boundary energy to converge, i.e., in the input script below, a distance of 12 nm was used. Figure 2. This is a plot of the accessibility of each grain boundary structure for an asymmetric tilt grain boundary. For complex boundaries, a large number of boundaries may need to be sampled to find the "global" minimum energy boundary.
In many cases, at the atomistic level, the minimum energy structure cannot be found by merely adjoining the two crystal lattices in the simulation cell. The relative translation of the two crystal lattices with respect to each other is important as well as the number of atoms within the grain boundary region. Therefore, to find the minimum energy structure, a large number of configurations with different relative translations is vital to finding the stable minimum energy structure and not just a metastable grain boundary structure. Figure 2 shows an example for an asymmetric tilt grain boundary, where the minimum energy structure was only found (accessed) 8.76% of the time with various translations. In more complex boundaries, the accessibility may be less than 0.1%.
The following input script shows how multiple translations and an atom deletion criteria are used to calculate the minimum energy structure. This input script for LAMMPS[3] can be called with a command of the form, "lmp_exe < input.script." This script contains loops over x-translations, z-translations, and atom overlap distances (an atom is deleted when an atom pair with a nearest neighbor distance is less than this distance). The unique minimum energy structures are saved as a dump file with the energy appended to the filename in a new folder specified by the 'gbname' variable. The dump files can then be easily scanned through for the global minimum energy structure.
|material model= none
|input deck=
# LAMMPS Input File for Grain Boundaries
# Mark Tschopp, Dec2009
# This file will generate numerous input files for LAMMPS
# using a large number of grain boundaries
# ---------- Setup Variables ---------------------
variable etol equal 1.0e-25
variable ftol equal 1.0e-25
variable maxiter equal 5000
variable maxeval equal 10000
variable latparam equal 2.855312
variable minimumenergy equal -4.122435
variable overlapboth equal 1
variable gbname index Fe_100STGB1
variable counter equal 0
variable inc equal "v_latparam / 6"
# Insert x,y,z sizes in LU and calculate in Angstroms
variable xsize1 equal "sqrt(0^2 + 2^2 + 1^2)"
variable zsize1 equal "sqrt(1^2 + 0^2 + 0^2)"
variable xsize2 equal "sqrt(0^2 + 2^2 + -1^2)"
variable zsize2 equal "sqrt(1^2 + 0^2 + 0^2)"
if ${xsize1} <= ${xsize2} then "variable xsize equal ${xsize1}" else "variable xsize equal ${xsize2}"
if ${zsize1} <= ${zsize2} then "variable zsize equal ${zsize1}" else "variable zsize equal ${zsize2}"
variable xlen equal "v_xsize * v_latparam"
variable zlen equal "v_zsize * v_latparam"
# Determine number of increments for displacement grid in the in-plane GB directions
variable xinc equal "floor(v_xlen / v_inc)"
variable zinc equal "floor(v_zlen / v_inc)"
# Implement overlap criterion
variable overlapinc equal 86
# ---------- Define loops for simulation ---------------------
label loopa
variable a loop ${xinc}
variable tx equal "(v_a-1) / v_xinc * v_xsize"
label loopb
variable b loop ${zinc}
variable tz equal "(v_b-1) / v_zinc * v_zsize"
label loopd
variable d loop ${overlapboth}
label loopc
variable c loop ${overlapinc}
variable overlapdist equal "0.275 + 0.005 * (v_c-1)"
# ---------- Calculate counter and create data directory ---------------------
variable ctemp equal ${counter}+1
variable counter equal ${ctemp}
variable ctemp delete
print "Counter: ${counter}"
shell mkdir ${gbname}
# ---------- Initialize Simulation ---------------------
clear
units metal
dimension 3
boundary p p p
atom_style atomic
# ---------- Create Atomistic Structure ---------------------
lattice bcc ${latparam}
region whole block 0.000000 6.384672 -121.308763 121.308763 0.000000 2.855312 units box
create_box 2 whole
region upper block INF INF 0.000000 121.308763 INF INF units box
lattice bcc ${latparam} orient x 0 2 1 orient y 0 -1 2 orient z 1 0 0
create_atoms 1 region upper
region lower block INF INF -121.308763 0.000000 INF INF units box
lattice bcc ${latparam} orient x 0 2 -1 orient y 0 1 2 orient z 1 0 0
create_atoms 2 region lower
group upper type 1
group lower type 2
# ---------- Define Interatomic Potential ---------------------
pair_style eam/fs
pair_coeff * * /cavs/cmd/data1/users/mtschopp/LAMMPS/lammps-12Nov09/potentials/Fe_2.eam.fs Fe Fe
neighbor 2.0 bin
neigh_modify delay 10 check yes
# ---------- Displace atoms and delete overlapping atoms ---------------------
displace_atoms upper move ${tx} 0 ${tz} units lattice
if $d == 1 then "delete_atoms overlap ${overlapdist} lower upper"
if $d == 2 then "delete_atoms overlap ${overlapdist} upper lower"
if $c == 1 then "variable atomprev equal 1"
variable natoms equal "count(all)"
print "Previous: ${atomprev}, Present: ${natoms}"
if ${atomprev} == ${natoms} then "jump GB_Fe_100STGB1.in loopend"
# ---------- Define Settings ---------------------
compute csym all centro/atom
compute eng all pe/atom
compute eatoms all reduce sum c_eng
# ---------- Run Minimization ---------------------
reset_timestep 0
thermo 10
thermo_style custom step pe lx ly lz press pxx pyy pzz c_eatoms
min_style cg
minimize ${etol} ${ftol} ${maxiter} ${maxeval}
# ---------- Run Minimization 2---------------------
# Now allow the box to expand/contract perpendicular to the grain boundary
reset_timestep 0
thermo 10
thermo_style custom step pe lx ly lz press pxx pyy pzz c_eatoms
fix 1 all box/relax aniso NULL 0.0 NULL vmax 0.001
min_style cg
minimize ${etol} ${ftol} ${maxiter} ${maxeval}
# ---------- Calculate GB Energy ---------------------
variable esum equal "v_minimumenergy * count(all)"
variable xseng equal "c_eatoms - (v_minimumenergy * count(all))"
variable gbarea equal "lx * lz * 2"
variable gbe equal "(c_eatoms - (v_minimumenergy * count(all)))/v_gbarea"
variable gbemJm2 equal ${gbe}*16021.7733
variable gbernd equal round(${gbemJm2})
print "After third minimization:"
print "GB energy is ${gbemJm2} mJ/m^2"
# Store number of atoms for overlap criterion, i.e., do not rerun equivalent configurations
variable atomprev equal "v_natoms"
# ---------- Dump data into Data file -------------
shell cd Fe_100STGB1
reset_timestep 0
timestep 0.001
velocity all create 20 95812384
fix 2 all npt 1 1 100 xyz 0 0 100 drag 0.2
dump 1 all custom 1000 dump.${counter}_${gbernd} id type x y z c_csym c_eng
run 0
shell cd ..
# ---------- End of loop structure -------------
label loopend
next c
jump GB_Fe_100STGB1.in loopc
variable c delete
next d
jump GB_Fe_100STGB1.in loopd
variable d delete
next b
jump GB_Fe_100STGB1.in loopb
variable b delete
next a
jump GB_Fe_100STGB1.in loopa
print "All done"
|results=
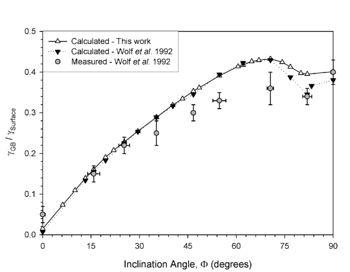
This methodology has resulted in structures that agree with HRTEM images and grain boundary energies that agree with experimentally-measured grain boundary energies. Figure 3 shows an example plot of the grain boundary energy versus inclination angle for a complex grain boundaries in Cu (Sigma 3 asymmetric grain boundaries). This system of boundaries displays a phase transformation at the boundary to the orthorhombic 9R phase for inclination angles of 70-90 degrees. The methodology used above agrees nicely with experimental results and previous atomistic calculations of Wolf and coworkers.
|acknowledgement=M.A. Tschopp would like to acknowledge funding provided under an NSF graduate fellowship for the initial work. Continued funding for investigating structure-property relationships in grain boundaries under the NEAMS (Nuclear Energy Advanced Modeling and Simulation) program is also acknowledged.
|references=The initial methodology was used in the following papers: